Directed Network (Class)#
The phylozoo.core.network.dnetwork module defines the
DirectedPhyNetwork class, the central representation of rooted, directed
phylogenetic networks in PhyloZoo.
A directed phylogenetic network is a graph with a distinguished root and leaves representing taxa. The directed edges encode ancestor–descendant relationships, with nodes of higher in-degree represent reticulate evolutionary events such as hybridization, recombination, or horizontal gene transfer.
All classes and functions on this page can be imported from the core network module:
from phylozoo import DirectedPhyNetwork
# or directly
from phylozoo.core.network.dnetwork import DirectedPhyNetwork
What is a directed phylogenetic network?#
Mathematically, a directed phylogenetic network is a weakly connected, directed acyclic multigraph (i.e. allowing parallel edges) with the following nodes:
Root: exactly one node with in-degree 0
Leaves: nodes with out-degree 0 and in-degree 1, each corresponding to a taxon
Tree nodes: internal nodes with in-degree 1 and out-degree ≥ 2
Hybrid nodes: internal nodes with in-degree ≥ 2 and out-degree 1
Internal nodes may be unlabeled. Leaf nodes carry labels (taxa), either provided explicitly or generated automatically. For technical completeness, the empty network and the single-node network (where root and leaf coincide) are also considered valid.
Example Networks
A simple directed tree, in which all internal nodes are tree nodes:
┌─A
┌───┤
│ └─B
────┤
│ ┌─C
└───┤
└─D
A directed network containing a hybrid node with two incoming edges:
┌────A
┌───┤
│ └──┬─B
| │
────┤ ├─C
| │
│ ┌──┴─D
└───┤
└────E
Creating a DirectedPhyNetwork#
A DirectedPhyNetwork is defined by its graph structure and optional attributes.
There are three main ways to construct such a network.
From edge and node specifications#
Edges may be given as tuples or dictionaries; nodes can explicitly be specified via (node_id, attributes) pairs, but are also automatically generated from the edges if not provided.
from phylozoo import DirectedPhyNetwork
network = DirectedPhyNetwork(
edges=[("root", "A"), ("root", "B"), ("root", "C")],
)
# or
network = DirectedPhyNetwork(
edges=[("root", "A"), ("root", "B"), ("root", "C")],
nodes=[("A", {"label": "A"}), ("B", {"label": "B"}), ("C", {"label": "C"})]
)
From an existing graph#
Directed networks can also be constructed from existing graph objects using the
dnetwork_from_graph() function. This is particularly useful when implementing algorithms that
operate on mutable graph representations.
from phylozoo.core.network.dnetwork.conversions import dnetwork_from_graph
import networkx as nx
G = nx.MultiDiGraph()
G.add_edge("root", "A")
network = dnetwork_from_graph(G)
Tip
The dnetwork_from_graph function is especially useful when writing your own algorithms.
It allows you to work directly with the underlying NetworkX or PhyloZoo graph object
of a DirectedPhyNetwork, after which you can convert back to a DirectedPhyNetwork at the
end of the algorithm.
# Example of writing an algorithm that works with the underlying graph
def my_algorithm(network: DirectedPhyNetwork):
# Get the underlying PhyloZoo DirectedMultiGraph
phylzoo_dm_graph = network._graph
# Get the underlying NetworkX MultiDiGraph
nx_graph = phylzoo_dm_graph._graph
# Do something with the NetworkX graph or the PhyloZoo DirectedMultiGraph
...
# Convert back to a DirectedPhyNetwork
return dnetwork_from_graph(nx_graph)
File Input/Output#
DirectedPhyNetwork supports reading and writing in standard formats:
eNewick (default): Extended Newick for rooted networks with hybrid nodes and edge attributes — see eNewick
DOT: GraphViz format for visualization and storage — see DOT format
# Load from file (auto-detects format by extension)
network = DirectedPhyNetwork.load("network.enewick")
# Load with explicit format
network = DirectedPhyNetwork.load("network.dot", format="dot")
# Load from string
network = DirectedPhyNetwork.from_string("(A,B);", format="enewick")
# Save to file
network.save("output.enewick")
network.save("output.dot", format="dot")
See also
The DirectedPhyNetwork class uses the
IOMixin interface, providing consistent file handling across PhyloZoo
classes. For details on the I/O system, see the I/O manual.
Validation on Construction#
Every DirectedPhyNetwork is validated at construction time using the validate() method. Validation
guarantees that the object represents a well-defined phylogenetic network:
the graph is acyclic and weakly connected,
there is exactly one root,
node degrees match the root / leaf / tree / hybrid definitions,
special attributes lie in their permitted ranges (see below for more details),
taxon labels are unique.
By default, invalid networks cannot be constructed. Validation can be disabled for performance-critical operations or when working with intermediate network states that may temporarily violate validation rules. See the Validation documentation for details on how to disable validation.
Attributes#
Nodes, edges, and the network itself may carry arbitrary attributes. Most attributes are stored without interpretation; however, the following attributes have a special, validated semantic role in directed phylogenetic networks.
Setting Attributes#
Attributes are set when creating a network by providing them in the edge, node, or attributes specifications:
Edge Attributes
Edge attributes are specified in edge dictionaries:
network = DirectedPhyNetwork(
edges=[
{"u": "root", "v": "A", "branch_length": 0.5, "bootstrap": 0.95},
{"u": "root", "v": "B", "branch_length": 0.3}
],
nodes=[("A", {"label": "A"}), ("B", {"label": "B"})]
)
Node Attributes
Node attributes are specified in node tuples:
network = DirectedPhyNetwork(
edges=[("root", "A"), ("root", "B")],
nodes=[
("A", {"label": "A", "custom_attr": "value1"}),
("B", {"label": "B", "custom_attr": "value2"})
]
)
Network Attributes
Network-level attributes are specified via the attributes parameter:
network = DirectedPhyNetwork(
edges=[("root", "A")],
nodes=[("A", {"label": "A"})],
attributes={"source": "simulation", "version": "1.0"}
)
Special edge attributes#
There are three special edge attributes that have a special, validated semantic role in directed phylogenetic networks and which may be used by certain algorithms.
branch_length (float)
Represents evolutionary distance along an edge.
bootstrap (float in [0, 1])
A statistical support value associated with the edge, for example from bootstrap or posterior analyses.
gamma (float in [0, 1])
The inheritance probability for hybrid edges (edges entering a hybrid node). For each hybrid node, the incoming gamma values must sum to 1.0.
These attributes can be accessed via dedicated methods:
network.get_branch_length("root", "A")
network.get_bootstrap("root", "A")
network.get_gamma("u1", "h")
Special node attribute#
There is one special node attribute that has a special, validated semantic role in directed phylogenetic networks and which may be used by certain algorithms.
label (string)
A unique identifier for a node. Labels must be unique across all nodes and must be strings. Leaf nodes always have labels (these are referred to as taxa), but internal nodes may be unlabeled. If a leaf node is created without an explicit label, one is automatically generated from the node ID.
Labels can be accessed via dedicated methods:
# Get label from node ID
label = network.get_label(node_id) # Returns label string or None
# Get node ID from label
node_id = network.get_node_id("A") # Returns node ID or None
Additional attributes#
Any additional node or edge attributes may be attached freely. Attributes without a special semantic role are preserved by the network object and by I/O operations but are not interpreted or validated. You can access these using generic attribute access methods:
# Get node attribute
node_attr = network.get_node_attribute(node_id, "custom_attr")
# Get edge attribute
edge_attr = network.get_edge_attribute(u, v, key=0, attr="custom_attr")
# Get network-level attribute
net_attr = network.get_network_attribute("source")
Accessing Network Properties#
The DirectedPhyNetwork class provides read-only access to fundamental structural
properties. These properties are computed from the validated network structure and cannot be
modified directly. All properties are cached for efficient repeated access.
Basic Properties#
Basic properties provide fundamental information about the network size and structure.
Nodes
The nodes cached property exposes a view of all node IDs (same interface as the
underlying graph). Use network.nodes() to iterate, or network.nodes(data=True) for
attributes.
# Get nodes (cached view)
list(network.nodes) # or network.nodes()
list(network.nodes(data=True)) # with attributes
Edges
The edges cached property exposes a view of all directed edges (same interface as the
underlying graph). Use network.edges(), network.edges(keys=True), or
network.edges(keys=True, data=True).
# Get edges (cached view)
list(network.edges) # or network.edges(keys=True)
list(network.edges(keys=True, data=True)) # with keys and attributes
Node and Edge Counts
The number_of_nodes() and number_of_edges() methods return the total count of
nodes and edges in the network, respectively.
# Get counts
num_nodes = network.number_of_nodes() # Total number of nodes
num_edges = network.number_of_edges() # Total number of edges
Check if Edge Exists
The has_edge() method checks if an edge exists between two nodes.
# Check if edge exists between nodes u and v
has_edge = network.has_edge(u, v) # Returns True if edge exists, False otherwise
# Check if edge exists with a specific key
has_edge_with_key = network.has_edge(u, v, key=0) # Returns True if edge exists with key 0, False otherwise
Node Types#
Network structure properties identify key nodes and their roles in the network topology.
Root
The root_node property returns the unique root node (in-degree 0).
# Get root node
root = network.root_node # Returns the root node ID
Leaves and Taxa
The leaves property returns a set of all leaf node IDs (out-degree 0).
The taxa property returns a set of all taxon labels (strings associated with leaves).
# Get leaves
leaves = network.leaves # Set of leaf node IDs
# Get taxa
taxa = network.taxa # Set of taxon labels
Internal Nodes
The internal_nodes property returns a set of all internal (non-leaf) node IDs.
# Get internal nodes
internal = network.internal_nodes # Set of internal node IDs
Tree and Hybrid Nodes
The tree_nodes property returns a set of all tree node IDs (internal nodes with in-degree 1 and out-degree ≥ 2).
The hybrid_nodes property returns a set of all hybrid node IDs (internal nodes with in-degree ≥ 2 and out-degree 1).
# Get tree nodes
tree = network.tree_nodes # Set of tree node IDs
# Get hybrid nodes
hybrid = network.hybrid_nodes # Set of hybrid node IDs
Edge Types#
Edge type properties classify edges according to their role in the network structure.
Tree and Hybrid Edges
The tree_edges property returns all tree edges (edges that are not hybrid edges)
as a set of (u, v, key) tuples. The hybrid_edges property returns all hybrid edges
(edges entering hybrid nodes) as a set of (u, v, key) tuples.
# Get edge type sets
tree_edges = network.tree_edges # Set of (u, v, key) tuples for tree edges
hybrid_edges = network.hybrid_edges # Set of (u, v, key) tuples for hybrid edges
# Example: iterate over hybrid edges
for u, v, key in network.hybrid_edges:
gamma = network.get_gamma(u, v, key=key)
print(f"Edge ({u}, {v}, {key}) has gamma={gamma}")
Node Connectivity#
Connectivity properties provide information about how nodes are connected in the network.
Node Degrees
The degree(), indegree(), and outdegree() methods return the total degree,
in-degree, and out-degree of a node, respectively.
# Get node degrees
total_degree = network.degree(v) # Total degree (in + out)
in_degree = network.indegree(v) # Number of incoming edges
out_degree = network.outdegree(v) # Number of outgoing edges
Neighbors
The parents(), children(), and neighbors() methods return iterators over
parent nodes, child nodes, and all adjacent nodes, respectively.
# Get parent nodes (nodes with edges pointing to v)
parents = list(network.parents(v)) # List of parent node IDs
# Get child nodes (nodes with edges from v)
children = list(network.children(v)) # List of child node IDs
# Get all neighbors (parents and children)
neighbors = list(network.neighbors(v)) # List of all adjacent node IDs
# Example: iterate over children
for child in network.children(v):
print(f"Node {v} has child {child}")
Incident Edges
The incident_parent_edges() and incident_child_edges() methods return iterators
over edges entering and leaving a node, respectively. These methods can optionally include
edge keys and attribute data.
# Get incoming edges (from parents)
incoming = list(network.incident_parent_edges(v))
# Returns: [(u1, v), (u2, v), ...]
# Get incoming edges with keys
incoming_with_keys = list(network.incident_parent_edges(v, keys=True))
# Returns: [(u1, v, key1), (u2, v, key2), ...]
# Get incoming edges with data
incoming_with_data = list(network.incident_parent_edges(v, data=True))
# Returns: [(u1, v, {'branch_length': 0.5, ...}), ...]
# Get outgoing edges (to children)
outgoing = list(network.incident_child_edges(v))
# Returns: [(v, w1), (v, w2), ...]
# Example: get all edge attributes for incoming edges
for u, v, key, data in network.incident_parent_edges(v, keys=True, data=True):
branch_length = data.get('branch_length')
bootstrap = data.get('bootstrap')
print(f"Edge ({u}, {v}, {key}): bl={branch_length}, bs={bootstrap}")
Copying#
The copy() method creates a shallow copy of the network. The copy shares the same
underlying graph structure but is a separate instance.
# Create a copy
network_copy = network.copy()
# Copies are independent instances
assert network_copy is not network
assert network_copy.root_node == network.root_node
See also
Some less commonly used or computationally expensive properties are available as functions in the advanced features module. These include blobs, omnian nodes, cut edges/vertices, and various network classifications. See Directed Network (Advanced) for details.
Visualization#
DirectedPhyNetwork can be visualized using the PhyloZoo visualization module. The visualization system supports multiple layout algorithms and styling options.
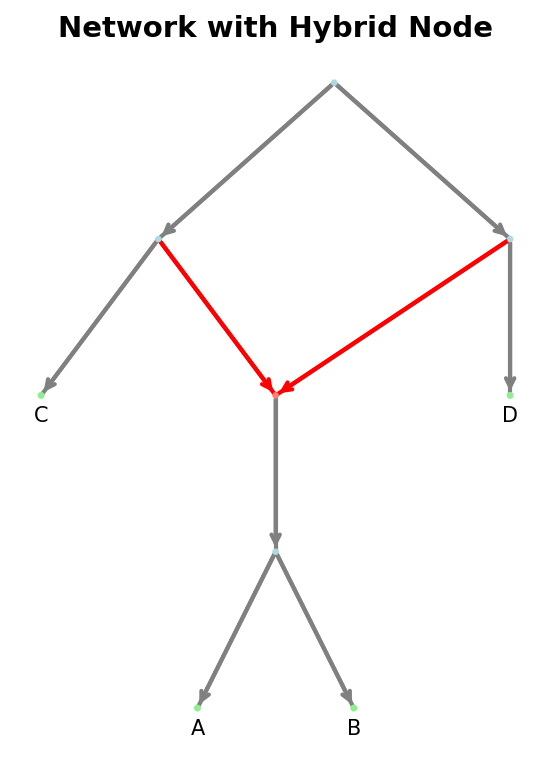
Example visualization of a directed network with a hybrid node.#
from phylozoo.viz import plot
# Plot network with default rectangular layout
plot(network)
For more visualization options, including different layout types, styling, and customization, see the Visualization documentation.
See Also#
API Reference - Complete function signatures and detailed examples
Directed Network (Advanced) - Advanced features, transformations, and classifications
Directed Generator - Level-k network generators
Semi-Directed Networks - Semi-directed network representations
I/O - File I/O operations and formats
Validation - Validation system and disabling validation
Visualization - Network visualization and plotting